3/10/2017. Text by Marta Daniela Santos.
One of the dangers of environmental changes at a global scale is the loss of biodiversity. A key questions is: are natural populations able to cope with these changes, adapting but maintaining their differences? A new study (*) led by Sofia Seabra, researcher at cE3c – Centre for Ecology, Evolution and Environmental Changes, based at the Faculty of Sciences of the University of Lisbon (Portugal) shows that natural selection can overcome the effects of history while maintaining the “genetic footprint”, thus preserving biodiversity.
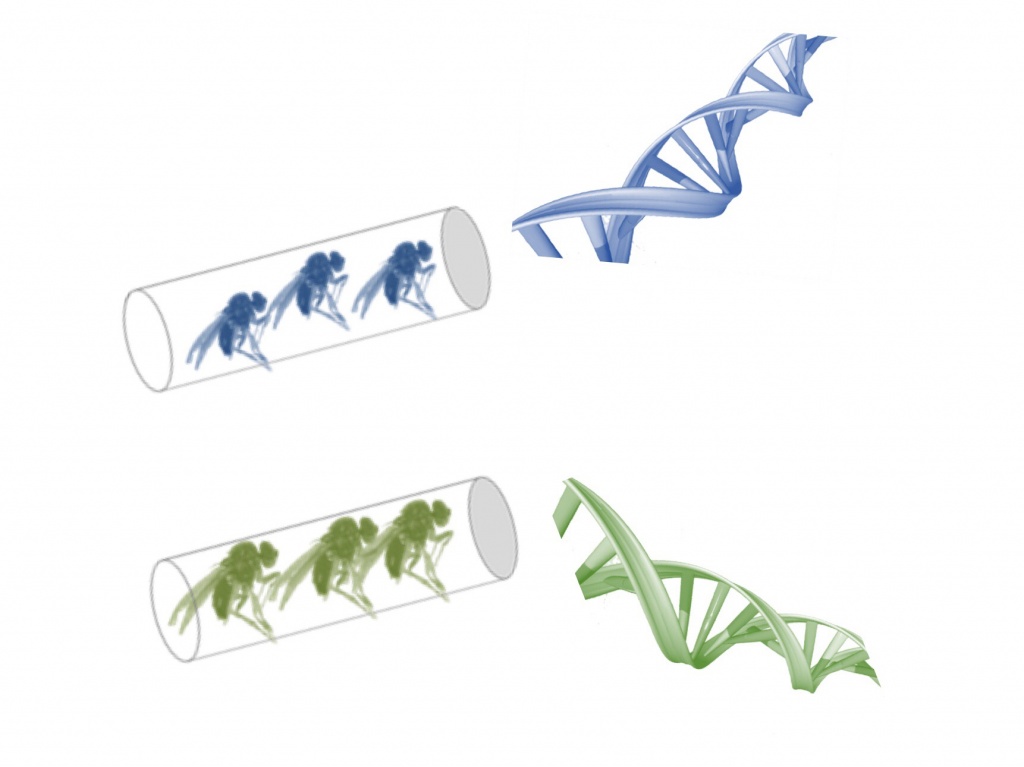
To understand how different populations of the same species respond to an identical selection pressure, researchers collected fruit flies of the species Drosophila subobscura in geographically distinct locations: Adraga (Portugal) and Groningen (Netherlands). Because they were adapted to different environments these populations differed, for example, in body size and physiological and reproductive characteristics. In addition, these populations differed in their evolutionary history: although they belong to the same species, each population has a genetic content that reflects how they have adapted to their specific environments over many generations.
Next, the researchers followed the evolution of these two populations in a new environment: the laboratory, where nutrients and temperature were controlled experimentally, among other conditions. After several generations, the two populations converged to similar adaptive solutions: flies from Adraga increased slightly in size approaching the size of the flies from Groningen, for example… What surprised the researchers was that, through genetic studies, they found that the two populations followed different genetic “routes” to reach these similar adaptive solutions – and the genetic “footprint” that distinguished them, showing that they belonged to two distinct populations, remained. That is, although they became similar from the point of view of their phenotype, from a genetic point of view they continued to present differences that showed that they belonged to different populations.
“Experimental evolution studies like this one allow us to elucidate the dynamics of adaptation to new environments, which is very difficult to do in studies done directly in nature. We have shown the complexity of molecular adaptation and the importance of taking into account the different biological levels that the adaptive evolution process involves”, explains Sofia Seabra, first author of the study and researcher at cE3c – Centre for Ecology, Evolution and Environmental Changes, based at the Faculty of Sciences of the University of Lisbon (Portugal).
This study also contributes with the publication of the first reference genome of the species Drosophila subobscura, an essential resource for genomic analyses that allowed researchers to demonstrate that the genetic “routes” followed by each population were different. As in the expression “All roads lead to Rome”: both populations arrived to Rome, but taking different roads.
The fact that the populations maintained their genetic differentiation after reaching similar adaptive solutions may indicate that they will have different adaptive potential to face future environmental changes. “The adaptive potential of populations will depend largely on the quantity and quality of available genetic variation. In this case, both populations had sufficient genetic variability to adapt, by different routes, to a relatively benign environmental change. If we were dealing with populations with low genetic variability to adapt to sudden changes in their environment, such as those caused by Man by introducing contaminants or degrading habitats, the answer could be very different”, explains Sofia Seabra.
(*) S.G.Seabra, I. Fragata, M.A. Antunes, G.S. Faria, M.A.Santos, V.C. Sousa, P. Simões, M.Matos, Different genomic changes underlie adaptive evolution in populations of contrasting history, Molecular Biology and Evolution. https://doi.org/10.1093/molbev/msx247
Tags: EE Cobig2